TissueWand, a rapid histopathology annotation tool
JPI (2020) · Paper
Abstract
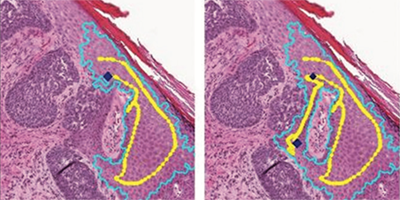
Background: Recent advancements in machine learning (ML) bring great possibilities for the development of tools to assist with diagnostic tasks within histopathology. However, these approaches typically require a large amount of ground truth training data in the form of image annotations made by human experts. As such annotation work is a very time-consuming task, there is a great need for tools that can assist in this process, saving time while not sacrificing annotation quality. Methods: In an iterative design process, we developed TissueWand – an interactive tool designed for efficient annotation of gigapixel-sized histopathological images, not being constrained to a predefined annotation task. Results: Several findings regarding appropriate interaction concepts were made, where a key design component was semi-automation based on rapid interaction feedback in a local region. In a user study, the resulting tool was shown to cause substantial speed-up compared to manual work while maintaining quality. Conclusions: The TissueWand tool shows promise to replace manual methods for early stages of dataset curation where no task-specific ML model yet exists to aid the effort.
Citation
@article{ 2020_jpi_tissuewand,
author = Martin Lindvall and Alexander Sanner and Fredrik Petré and Karin Lindman and Darren Treanor and Claes Lundström and lowgren,
title = "TissueWand, a rapid histopathology annotation tool",
year = 2020,
pages = 27,
doi = "10.4103/jpi.jpi_5_20",
}